Analysis and Comparison of the Impact of Statistical Feature Reduction for Classification using Machine Learning
DOI:
https://doi.org/10.65205/jcct.2026.e3002Keywords:
Disease Classification, Feature Selection, Hyperparameter, Machine LearningAbstract
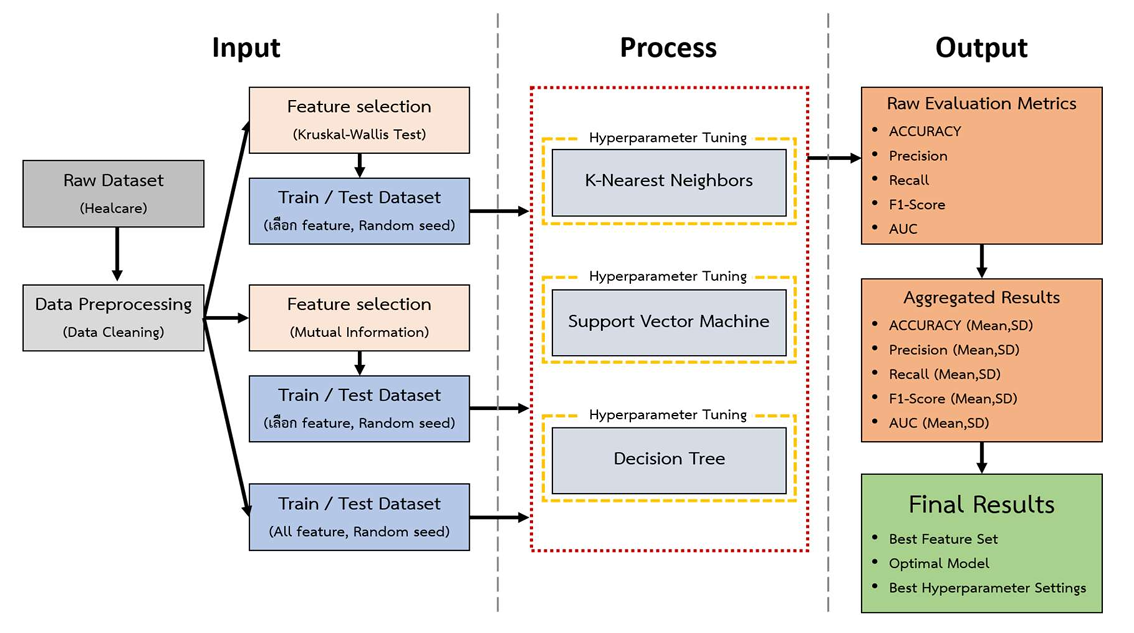
This study aims to analyze the impact of statistical feature reduction on classification performance and to identify the optimal hyperparameter configurations for predicting medical conditions. Three machine learning algorithms—Support Vector Machine (SVM), Decision Tree (DT) and K-Nearest Neighbors (KNN) were compared using the Healthcare Risk Factors Dataset. Two filter-based feature selection methods, the Kruskal–Wallis test (KW) and Mutual Information (MI), were employed to generate a full feature set and feature subsets at the 25th, 50th, and 75th percentiles. Model performance was evaluated using 5 repeats of 5-fold Stratified K-Fold Cross-Validation, integrated with systematic hyperparameter tuning. The experimental results demonstrated that SVM achieved the highest performance using the full feature set (C = 300), yielding an accuracy of 0.9214 ± 0.0005, precision of 0.9212 ± 0.0005, recall of 0.9214 ± 0.0005, and an F1-Score of 0.9210 ± 0.0005. This accuracy significantly outperformed DT (0.8752 ± 0.0028) and KNN (0.7916 ± 0.0015). Furthermore, reducing the feature set to the 75th percentile using the KW method maintained a high accuracy of 0.9165 ± 0.0006, representing a marginal decline of only 0.49%. These findings indicate that SVM is highly suitable for disease diagnosis within this dataset. Moreover, statistical feature reduction effectively minimizes computational complexity without a significant loss in accuracy.
Downloads
References
Ahmed, A. (2025). Healthcare Risk Factors Dataset [Dataset]. Kaggle. https://www.kaggle.com/datasets/abdallaahmed77/healthcare-risk-factors-dataset
Bashir, S., Khattak, I. U., Khan, A., Khan, F. H., Gani, A., & Shiraz, M. (2022). A Novel Feature Selection Method for Classification of Medical Data Using Filters, Wrappers, and Embedded Approaches. Complexity, 2022(1), 8190814. https://doi.org/10.1155/2022/8190814 DOI: https://doi.org/10.1155/2022/8190814
Clarke, R., Ressom, H. W., Wang, A., Xuan, J., Liu, M. C., Gehan, E. A., & Wang, Y. (2008). The Properties of High-Dimensional Data Spaces: Implications for Exploring Gene and Protein Expression Data. Nature Reviews Cancer, 8(1), 37-49. https://doi.org/10.1038/nrc2294 DOI: https://doi.org/10.1038/nrc2294
Guyon, I., & Elisseeff, A. (2003). An Introduction to Variable and Feature Selection. Journal of Machine Learning Research, 3, 1157-1182.
Jothi, N., Syed-Mohamed, S. M., & Rajagopal, H. (2022, July 20-22). Hybrid Feature Selection using Shapley Value and ReliefF for Medical Datasets. 2022 International Conference on Inventive Computation Technologies, 351-355. Institute of Electrical and Electronics Engineers. https://doi.org/10.1109/icict54344.2022.9850833 DOI: https://doi.org/10.1109/ICICT54344.2022.9850833
Kavakiotis, I., Tsave, O., Salifoglou, A., Maglaveras, N., Vlahavas, I., & Chouvarda, I. (2017). Machine Learning and Data Mining Methods in Diabetes Research. Computational and Structural Biotechnology Journal, 15, 104-116. https://doi.org/10.1016/j.csbj.2016.12.005 DOI: https://doi.org/10.1016/j.csbj.2016.12.005
Kruskal, W. H., & Wallis, W. A. (1952). Use of Ranks in One-Criterion Variance Analysis. Journal of the American Statistical Association, 47(260), 583-621. https://doi.org/10.1080/01621459.1952.10483441 DOI: https://doi.org/10.1080/01621459.1952.10483441
Majed, R. J., Al-Heddi, R. M., & Zeki, A. M. (2023, October 25-26). Parkinson’s Disease Detection Using Data Mining Models: A Comparative Study. 2023 4th International Conference on Data Analytics for Business and Industry, 290-294. Institute of Electrical and Electronics Engineers. https://doi.org/10.1109/icdabi60145.2023.10629302 DOI: https://doi.org/10.1109/ICDABI60145.2023.10629302
Narwane, S. V., & Sawarkar, S. D. (2021, August 27-29). Dimensionality Reduction of Unbalanced Datasets: Principal Component Analysis. 2021 Asian Conference on Innovation in Technology, 1-6. Institute of Electrical and Electronics Engineers. https://doi.org/10.1109/asiancon51346.2021.9544971 DOI: https://doi.org/10.1109/ASIANCON51346.2021.9544971
Prusty, S., Patnaik, S., & Dash, S. K. (2022). SKCV: Stratified K-Fold Cross-Validation on ML Classifiers for Predicting Cervical Cancer. Frontiers in Nanotechnology, 4, 972421. https://doi.org/10.3389/fnano.2022.972421 DOI: https://doi.org/10.3389/fnano.2022.972421
Remeseiro, B., & Bolon-Canedo, V. (2019). A Review of Feature Selection Methods in Medical Applications. Computers in Biology and Medicine, 112, 103375. https://doi.org/10.1016/j.compbiomed.2019.103375 DOI: https://doi.org/10.1016/j.compbiomed.2019.103375
Saha, P., Patikar, S., & Neogy, S. (2020, October 2-4). A Correlation-Sequential Forward Selection Based Feature Selection Method for Healthcare Data Analysis. 2020 IEEE International Conference on Computing, Power and Communication Technologies, 69-72. Institute of Electrical and Electronics Engineers. https://doi.org/10.1109/gucon48875.2020.9231205 DOI: https://doi.org/10.1109/GUCON48875.2020.9231205
Salehin, S., Islam, A. J., Iqbal, A. M., Barua, L., Islam, K., & Uddin, A. (2024, December 7-8). A Comparative Analysis of Feature Selection Methods on the Accuracy of Heart Disease Prediction Models. 2024 International Conference on Recent Progresses in Science, Engineering and Technology, 1-6. Institute of Electrical and Electronics Engineers. https://doi.org/10.1109/icrpset64863.2024.10955882 DOI: https://doi.org/10.1109/ICRPSET64863.2024.10955882
Shannon, C. E. (1948). A Mathematical Theory of Communication. Bell System Technical Journal, 27(3), 379-423. https://doi.org/10.1002/j.1538-7305.1948.tb01338.x DOI: https://doi.org/10.1002/j.1538-7305.1948.tb01338.x
Siburian, R. H., Km Nasution, M., & Tarigan, J. T. (2025, January 21). Optimization Of KNN, SVM, And SVM Kernel in Water Potability Prediction with Hyperparameter Approach. 2025 International Conference on Computer Sciences, Engineering, and Technology Innovation, 576-581. Institute of Electrical and Electronics Engineers. https://doi.org/10.1109/icocseti63724.2025.11019055 DOI: https://doi.org/10.1109/ICoCSETI63724.2025.11019055
Uddin, S., Khan, A., Hossain, M. E., & Moni, M. A. (2019). Comparing Different Supervised Machine Learning Algorithms for Disease Prediction. BMC Medical Informatics and Decision Making, 19(1), 281. https://doi.org/10.1186/s12911-019-1004-8 DOI: https://doi.org/10.1186/s12911-019-1004-8
Downloads
Published
How to Cite
Issue
Section
Categories
License
Copyright (c) 2026 Journal of Computer and Creative Technology

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.